Understanding our fish, keeping our seas sustainable
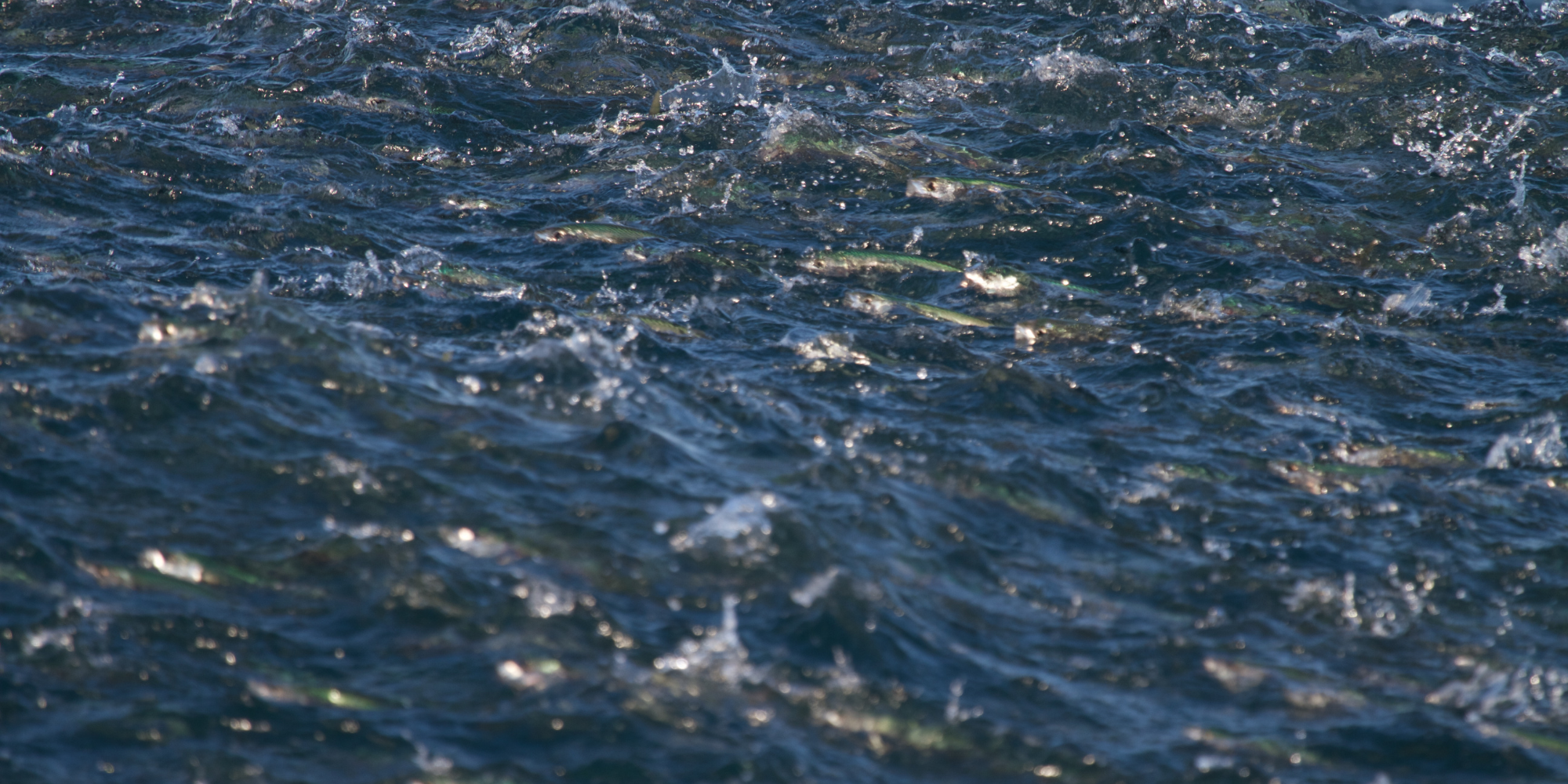
How many fish swim in the waters off our shores? Which species are out there, and where are they? Understanding and managing fish populations has been critical to fishers for centuries. When their knowledge isn’t good enough, disaster can strike – as happened when the Atlantic herring numbers in the North Sea collapsed towards the end of the twentieth century.
Genomics is now providing the tools to build upon generations of understanding of our seas, and draw a much more detailed picture of what happens beneath the waves. This is a key part of efforts to keep our ocean healthy.
The Darwin Tree of Life project has published its first Genome Note for a commercially fished oceanic species, the Atlantic horse mackerel (Trachurus trachurus). Scientists studying fish populations are already using our high-quality reference genomes to support their research.
For starters: Atlantic horse mackerel
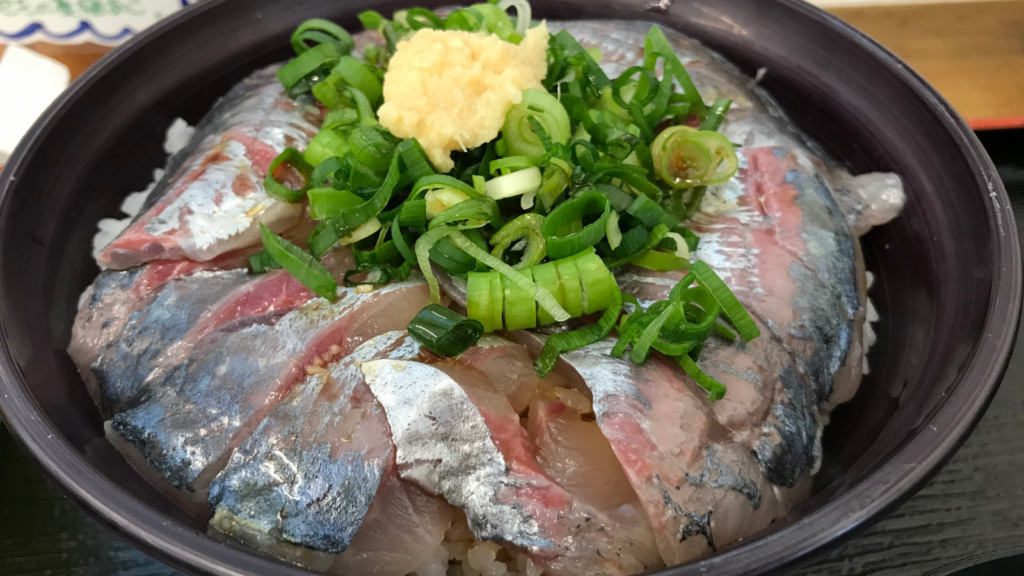

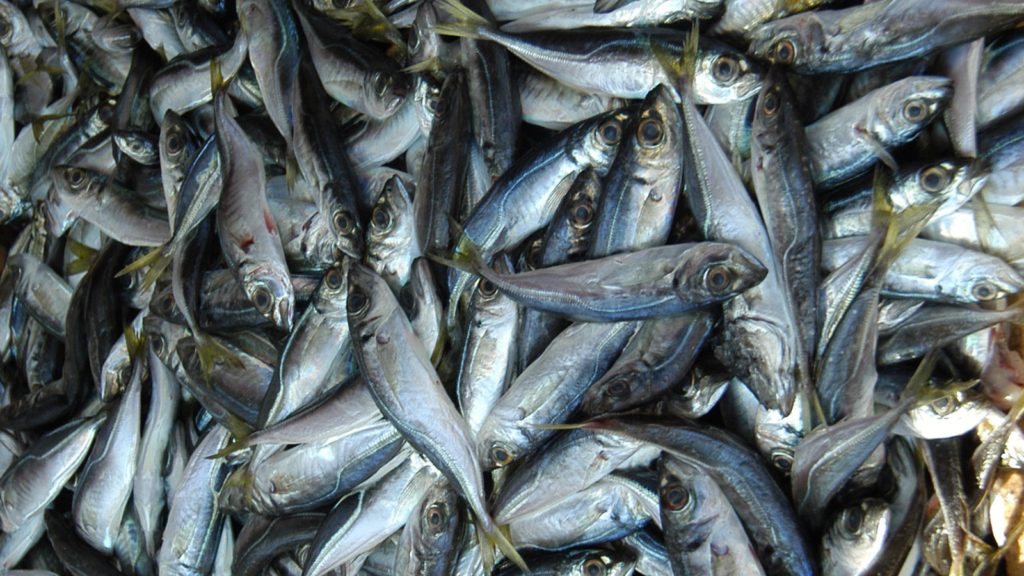
The Atlantic horse mackerel is found from Norway to Africa, along the Mediterranean and up to the Black Sea. Though seldom gracing northern European plates, it is caught in huge numbers and used in everything from sushi to fish meal – a product for feeding farmed animals, including other fish.
However, the northern stock of Atlantic horse mackerel has recently declined in number. It is now classified as “vulnerable” on the IUCN Red List.
One of the scientists using our new reference genome for Trachurus trachurus is Leif Andersson, professor in comparative genetics and functional genomics at Uppsala University, Sweden. Leif and his team use genomics to understand genetic variation and ecological adaptation of the horse mackerel throughout its large range.

“Genomics allows us to study every gene in a genome. It’s a complete revolution in how we study biology,” Leif explains.
In the field of population genomics, scientists study genetic markers – small sequences of DNA known as SNPs (pronounced “snips”) – to look for any differences between fish populations, for example between those living off the coast of Norway versus those west of Ireland. The findings then help inform how those populations – known as “stocks” – are managed by the fishing industry.
But to do this in any detail, researchers first need a full genome sequence as a reference to compare back to.
In their previous studies on Atlantic herring (Clupea harengus), Leif’s team assembled their own genome sequence. But this costs time and money.
“Now we contact people at the Darwin Tree of Life project,” says Angela Fuentes Pardo, who worked with Leif on both the herring and horse mackerel studies. “If they’re on the way to making an assembly of the species, then there’s no need for us to do that and we can focus on the population studies. For our horse mackerel studies that was exactly what happened.”
Taking stocks: towards sustainable fishing

What is a fish stock? It’s a tougher question than you might think. A stock is essentially a population of one species that is more or less isolated from other members of the same species, and therefore self-sustaining. So how do fishers (and scientists) know when they are dealing with one stock rather than another?
Historically, stocks were seen geographically. For example, if there was an abundance of fish in the seas off Norway and another hotspot in the Irish Sea, but nothing much in between, these would be assumed to be two distinctive stocks.
More recently, scientific studies have revealed a more complex picture. For example, fish move around a lot throughout their lifecycle – spawning in one part of the ocean and spending their adult lives elsewhere. Two different stocks might live as adults in the same area, but spawn in totally different places. Or vice versa.
And the picture is quite diverse for different species of fish, for example stock structures are quite different for Atlantic herring compared to Atlantic horse mackerel. Meanwhile every European eel (Anguilla anguilla) spawns at the same location in the Sargasso Sea, and therefore all European eels belong to the same stock.
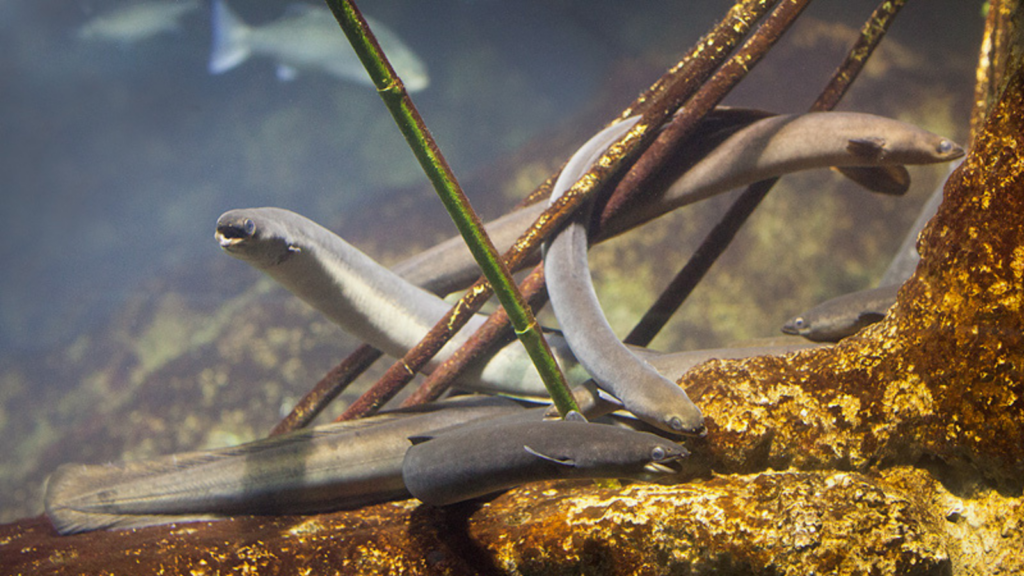

“My impression is that the fishing industry is very interested in these studies,” says Leif. “There’s this reputation that they just want to fish as much as possible, and of course they are commercial operators and want to make profit. But they are aware that the new tools of genomics are really valuable for them, to see which stocks they can utilise. And they don’t want to see stocks collapse because then their business is gone!”
Indeed, the Trachurus trachurus population studies carried out by Leif and his team were commissioned and funded by the Northern Pelagic Working Group, part of the European Association of Fish Producers Organisations.
A fish swam here: using environmental DNA
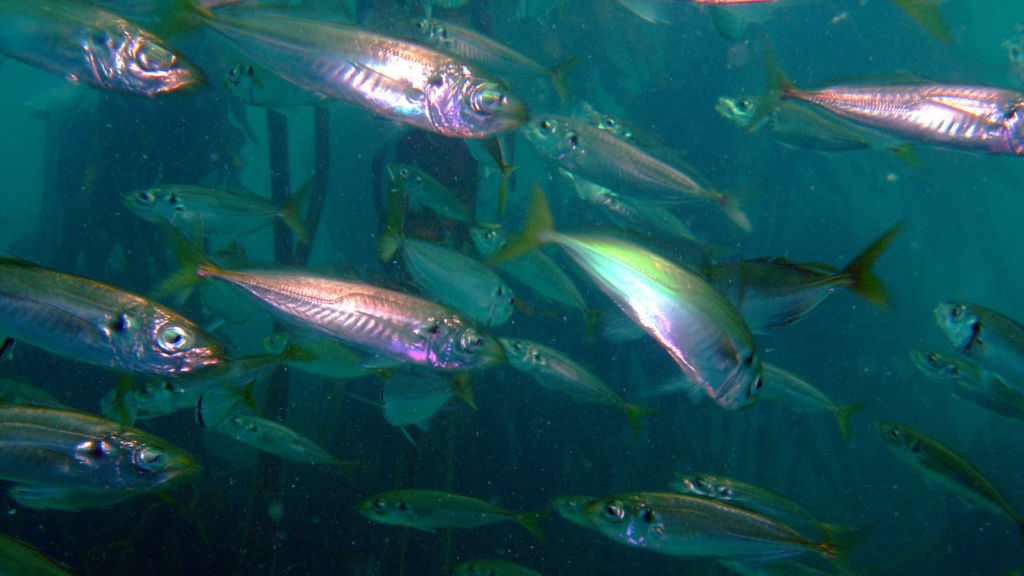

The specific mackerel that ended up in our genome sequencing pipeline was originally collected as part of an environmental DNA project called SeaDNA. It was fished out of Southampton Water, off the Hampshire coast, and fortuitously snap-frozen quickly enough to prevent the DNA from deteriorating.
Environmental DNA is another exciting development in genomics. This lets you take a non-living sample from the environment – for example soil, air or seawater – and test to see which species have passed through. One of the many benefits is this avoids destructive sampling practices, for example trawling the ocean floor or fishing in a protected area.
There is a “surprising amount” of genetic material in a sample of seawater, explains Martin Genner, the principal investigator on the SeaDNA project. But one of the first things the team noticed was a lack of existing genomic data for the species they wanted to study.
Genomics has the potential to answer all sorts of scientific questions. It allows researchers to not only study the occurrence and abundance of species today, but also how genes are flowing between different fish stocks over time. It may even answer mysterious questions about how new species diversity evolves in the sea – this is a longstanding head-scratcher, considering there are no obvious physical barriers to stop different populations breeding and ultimately forming new species.
“But before we can even go anywhere near those questions we first need high-quality reference genomes,” says Martin.
That is exactly what the Darwin Tree of Life project aims to provide, with the goal of producing high-quality, chromosomal-level reference genomes for every species in British and Irish waters.
Leif Andersson welcomes the initiative, but acknowledges this is only the beginning. “Darwin Tree of Life assemblies are a great resource, but they need to be complemented by population studies. So the hope is that funding will come for those studies also.”