Genomes great and small: The diversity of plants
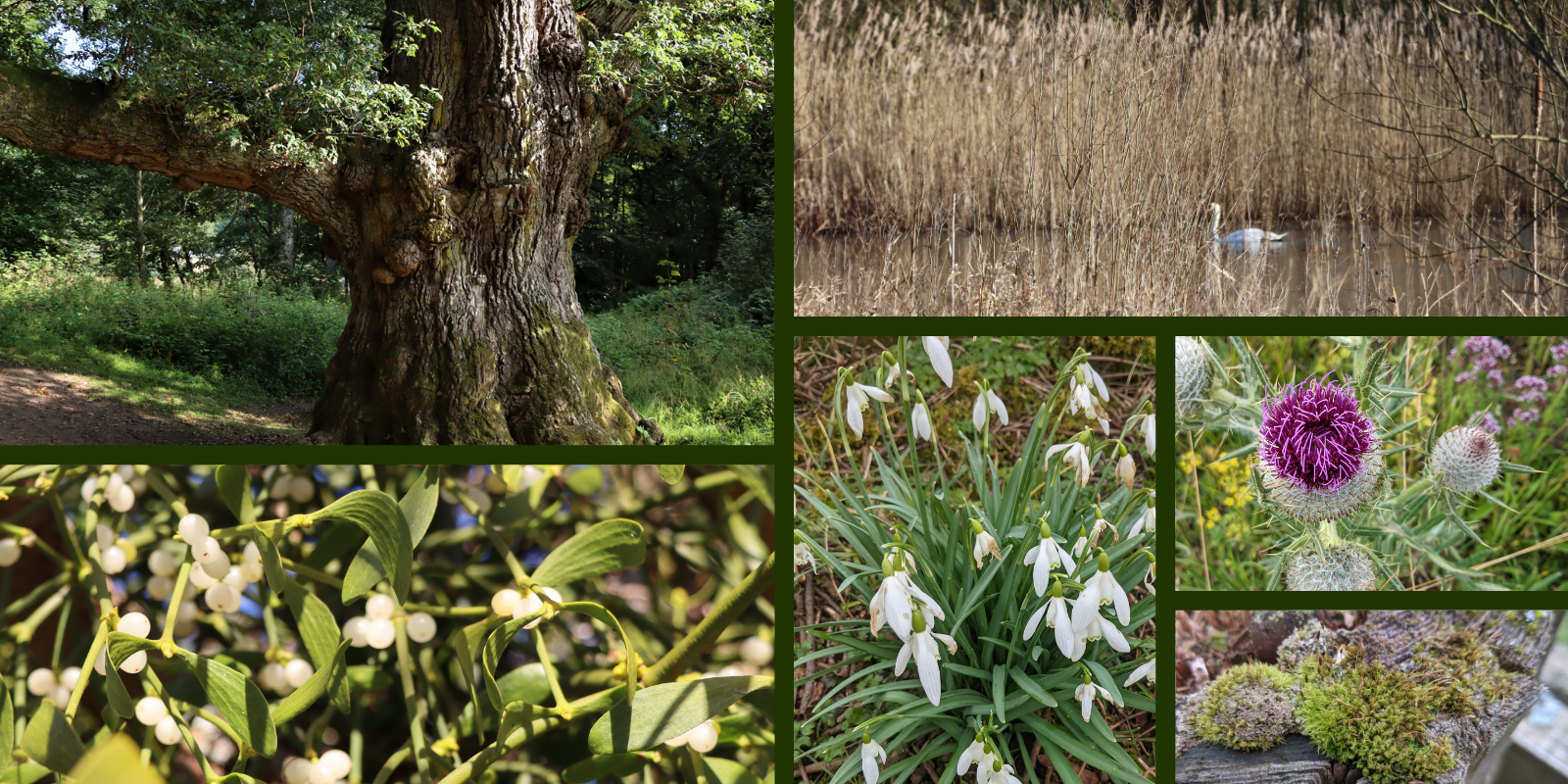
Plants are the most diverse group of organisms on the planet – genomically speaking. Paris japonica, the Japanese canopy plant, has the largest genome of any organism analysed to date. At 149 billion base pairs (giga bp or Gbp) of DNA, it’s about 50 times bigger than the human genome. At the other end of the scale, the flowering plant with the smallest genome is Genlisea tuberosa, a tiny carnivorous plant found in Brazil, coming in at 61 million base pairs (mega bp or Mbp).
Plants also have an extraordinarily large range of ‘ploidy’ – the number of complete sets of chromosomes in their cells. Humans and many animals are diploid, with two copies of each chromosome, one from each parent. Plant species may have anything from two to 96 copies of each chromosome. One 96-ploid species of fern has over 1,400 chromosomes per cell – the highest chromosome number known to science.
The often huge amounts of DNA inside plant cells affects how they function. It may also influence their ability to adapt and evolve, especially in periods of rapid environmental change. Understanding how plants evolve and survive in the face of climate change is crucial for the future, especially considering 90% of humanity’s energy intake comes from just 15 species of plants. Knowledge of plant genomes will help with agriculture and biotechnology; wild relatives of domesticated species may harbour traits that will help them adapt to, for example, global heating, nutrient loss, or aridification.

To date, just over 900 of the estimated 450,000 plant species on Earth have had a genome sequenced. Part of the reason is that their wide variety, and often huge amounts of DNA, make decoding plant genomes a complex task.
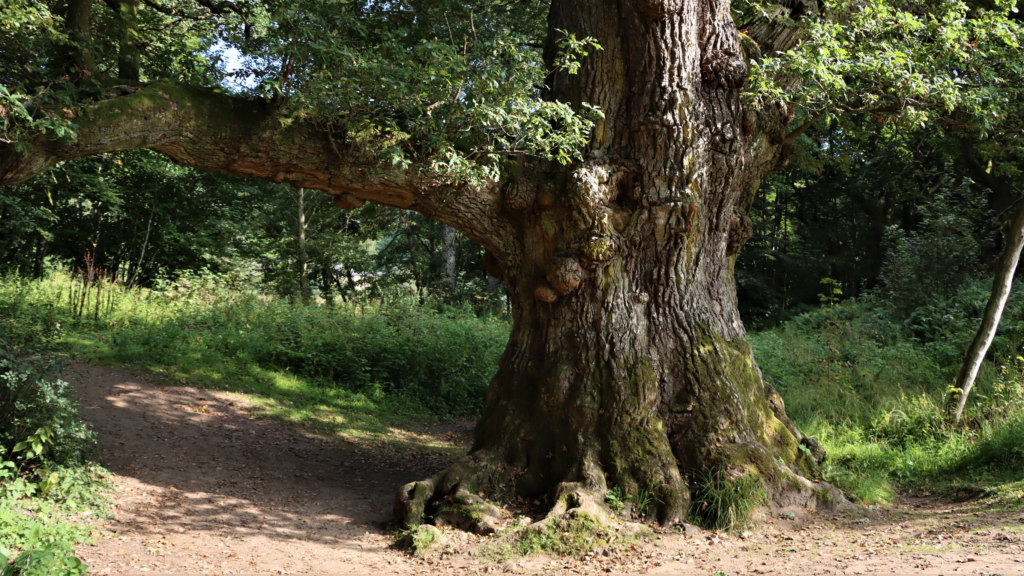
Researchers on the Darwin Tree of Life project are aiming to sequence the genomes of all complex life in Britain and Ireland, including plants. The project is about to publish its first plant genome – the common oak, Quercus robur. It has a small genome of about 800 Mbp – a third of the size of the human genome: if you stretched out all the DNA from a single cell of an oak tree, it would stretch to just 66cm. Our own genome would reach 2m, and Paris japonica would be 100m long.
| Species | Common name | Ploidy level | Genome size (Gbp/1C) | Approx. length of DNA in a cell |
| Ostreococcus tauri | a unicellular marine green alga | Haploid | 0.012 | 8 mm |
| Ananas comosus | Pineapple | Diploid | 0.5 | 35 cm |
| Fragaria x ananassa | Strawberry | Octoploid | 0.6 | 40 cm |
| Coffea arabica | Coffee | Tetraploid | 1.2 | 80 cm |
| Cocos nucifera | Coconut | Diploid | 2.7 | 1.8 m |
| Dionaea muscipula | Venus fly trap | Diploid | 2.8 | 1.9 m |
| Allium cepa | Onion | Diploid | 15.6 | 10.4 m |
| Aloe vera | Aloe | Diploid | 16 | 10.7 m |
| Triticum aestivum | Bread wheat | Hexaploid | 16.9 | 11.3 m |
| Pinus sylvestris | Pine tree | Diploid | 22.5 | 15.0 m |
| Galanthus nivalis | Snowdrop | Diploid | 35.3 | 23.5 m |
| Viscum album | Mistletoe | Diploid | 95.1 | 63.3 m |
| Paris japonica | Japanese canopy plant | Octoploid | 148.8 | 100 m |
Dr. Ilia Leitch is a plant biologist at the Royal Botanic Gardens Kew, where she researches the genomics of plants and plant evolution. The gardens, together with the Royal Botanic Garden Edinburgh and the Marine Biological Association based at Plymouth, are coordinating the collecting of a specimen of every single plant species in Britain and Ireland for DNA sequencing as part of the Darwin Tree of Life project.
Ilia is involved in overseeing the collection of plants for the project. Many UK species have been gathered from Kew’s extensive plant collections, although she also co-ordinates collecting trips for those that aren’t available at Kew, as well as growing up rare plants from seeds stored in the gardens’ Millennium Seed Bank.
For each species, four different samples must be collected – one sample is sent to the Sanger Institute for genome sequencing, and one to Edinburgh for DNA barcoding, to assist with species identification. Two samples are kept at Kew. Of these, one goes to the herbarium to provide a permanent physical specimen of each plant that is sequenced, while the other is sent to the laboratory to assess its genome size, data that are invaluable for teams at Sanger, so they know how much DNA they need to sequence.

“I’m interested in the evolutionary significance of genome size,” says Ilia, “because to some extent, it is baffling why one plant may have so much more DNA than another. I’m interested in understanding how such diversity evolved. What types of DNA sequences make up genomes of different sizes? How is that regulated? I’m also looking at the ecological consequences – does it matter if you’re a plant with a massive great genome growing and trying to compete for resources next to a plant that has got a tiny genome? Does that impact your ability to survive? I’m aiming to measure and understand the trade-offs.”
One of the main effects of genome size on a plant’s functions is on its ability to grow, and the type of lifecycle it adopts. For example, Arabidopsis – the most studied plant in the world – has a very small genome. It is an ephemeral plant, that can grow from seed, to plant, to seed, in just six to eight weeks. Plants with huge genomes can’t grow that quickly – they take a lot more time and energy to copy all their DNA every time they divide their cells. That means it takes longer to mature, and so plants with large genomes are restricted to being perennial, only being able to produce flowers after more than a year.
“Having this opportunity to study genome size and composition for a whole flora – it’s really exciting,” says Ilia. “We’re also interested in how genome size may impact, for example, the extinction risk of plants. We are keen to uncover the distribution of genome sizes across the UK, how that has varied over time, and the potential trajectory under environmental change. If we get a heatwave or the climate changes, those plants with the bigger genomes – the slower growing perennials – are probably going to be at greater risk of extinction.”

The material of evolution
Polyploidy, on the other hand, may be an evolutionary advantage for some plant species. Researchers have uncovered that whole genome duplications have occurred multiple times in most plant species. Initially this results in a species having duplicated copies of each chromosome and a larger genome, though often some of the extra DNA is whittled away and lost, to return to a more streamlined genome.
An extra copy of the genome results in more genetic diversity within a species, and also a wider variety of traits. Polyploidy can also lead to bigger cells, which may mean bigger seeds or fruit – something that has long been selected for by farmers and crop breeders. However, polyploidy may also lead to genetic instability and infertility. Polyploidy can also have ‘costs’, such as increased demand for nutrients such as nitrogen and phosphorous needed to build the genome. This may in turn also play a role in influencing a species competitive ability in the landscape, particularly those where nutrients are scarce.
“Having platinum-grade, chromosome level genome assemblies for all native plants in Britain and Ireland will mean that we can really start to probe into understanding what’s happening within genomes of different sizes. We know that repetitive DNA sequences contribute a lot to the differences in genome size between species. But now we can see where all the different types of repetitive DNA sequences are located within the genome and how their activity varies in genomes of different sizes or between species growing in different environments. In addition, we can see which repeat sequences are close to genes and if so, are they playing a role in regulating the gene expression. There are all sorts of molecular and ecological questions you can ask now which you just couldn’t do before. It is like uncovering a new landscape to explore,” says Ilia.

Challenges
While the variation in genome size and copy number is exciting, it does present challenges. For the bioinformaticians determining the genome sequences at the Sanger Institute, it means there is a lot of complexity in the data. Knowing the genome size in advance helps, as it can be used to check a new genome sequence is as expected, or if there are parts missing or duplicated in the data. Genome size information is fed through directly from Kew to Sanger.
Big genomes also mean big data. Dr. Max Brown is a postdoctoral scientist at the Sanger Institute researching the evolutionary history of some of the UK’s most loved plant species, including oak and apple trees.
“A major challenge of working on this project is the huge amount of data you have to crunch through. There’s so much data it can take a long time to get any results back. The algorithms that are being implemented are constantly improving, but you still need huge computational power. We are lucky at Sanger that we do have such resources, but it still takes time.”

Future research
Plants, whatever their genome size, underpin all aspects of our everyday lives, from the food we eat and the air we breathe, to medicines, clothes and buildings.
Plant sciences have a vital role in addressing the critical global challenges of climate change and food security. The Darwin Tree of Life team hopes that data from plant genome sequences will underpin future research into plant development, biodiversity and evolution, and will help those studying medicinally important compounds, biofuels and sustainable agriculture.
This article was adapted from an article written by Alison Cranage, science writer at the Wellcome Sanger Institute, and originally published on the Wellcome Sanger Institute blog.