DToL in 2022: First 500 genome assemblies as project creates a buzz
As of December 2022, the Darwin Tree of Life project has released 500 reference-quality genome assemblies to public databases, ready to be used by researchers across the planet. The project is also making its mark on the airwaves and in print, with articles, podcasts, radio interviews and even BBC national news coverage. The successes of this year have been down to the tireless DToL team – whether in the field or the lab, on computers or taking our science on the road.
Below are a selection of highlights chosen by some of the DToL partners.
Releasing our first 500 assemblies

For everyone at Sanger’s Tree of Life Programme, 2022 will be looked back on as the year our genome production pipeline really powered into action. This initial stage of the Darwin Tree of Life project has given proof of concept for this ambitious genomics venture. Could we create a series of scientific processes that take an organism in the wild and transform it into a top-quality, chromosomal-level genome assembly representing its entire species? Could those processes be repeated again and again for thousands of species spanning every branch of the tree of life? Could a network of partner organisations focused on ecology, informatics and analysis be brought together to achieve this goal?
The answer is a resounding yes! In the last 12 months we have more than doubled the number of DToL genome assemblies on public databases and tripled our catalogue of published Genome Notes.
The graphic above shows the diversity of our first 500 assemblies across the tree of life. There has undoubtedly been a bias towards arthropods, for a number of reasons. They are easy to collect and sequence – for example, moths fly towards light, arthropod DNA is relatively easy to extract in the lab, and many have smaller genomes. But we also wanted to showcase the project’s potential for aiding comparative genomics, with DToL scientists already publishing arthropod studies based on our genome assemblies.
There is also a significant breadth of diversity emerging. In 2022 we published Genome Notes for the first fungi, cnidarians, tunicates, molluscs and – most recently – plants. The first protists are due to be published soon. Getting to grips with this dazzling array of organisms and their genomes is a key achievement in a very successful year.
The first plant genomes are published
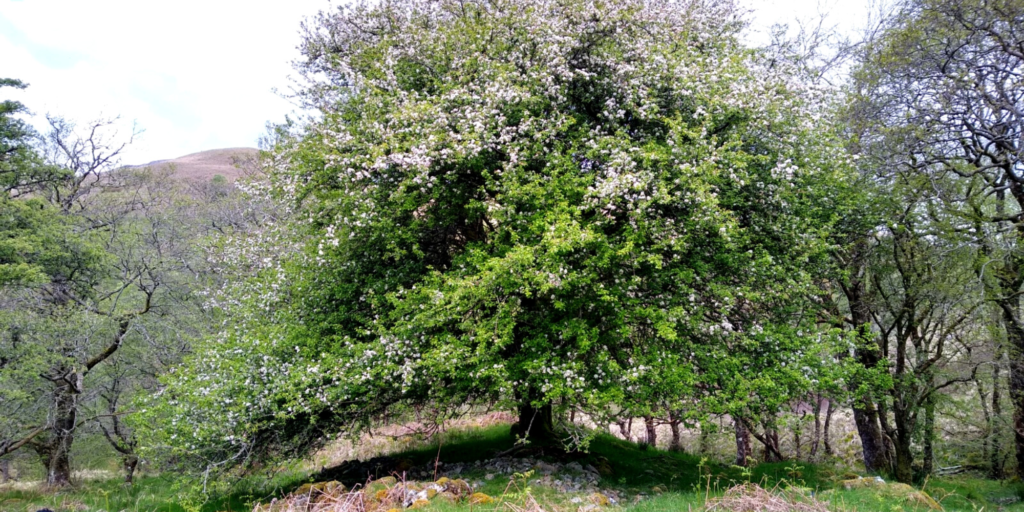

You wait ages to publish some reference genomes for plants, then five come at once! We sequenced Britain and Ireland’s only native wild apple (Malus sylvestris) plus four heritage cultivars of Malus domestica originally grown on these shores. This is just part of a deeper apple-based project our botanists at Edinburgh and Kew, bioinformaticians at Sanger, and other collaborators have been involved with since DToL began, also producing short-read DNA sequences to compare over 40 other varieties of locally grown apples.
These genomes can help answer questions about our apple history, the species’ evolution, and how to protect the precariously positioned crab apple. Its genetic integrity is being undermined by hybridisation with widely-planted domestic relatives. Nearly 30% of the ‘wild’ apple trees surveyed in a recent study in northern Britain turned out to be of hybrid origin.
Learn more about our apple genomes here. And if you fancy mulling over some genomics with a hot cider this winter, check out the short Scider videos we made looking at the science of this apple-based beverage.
Mistletoe: Britain & Ireland’s largest genome
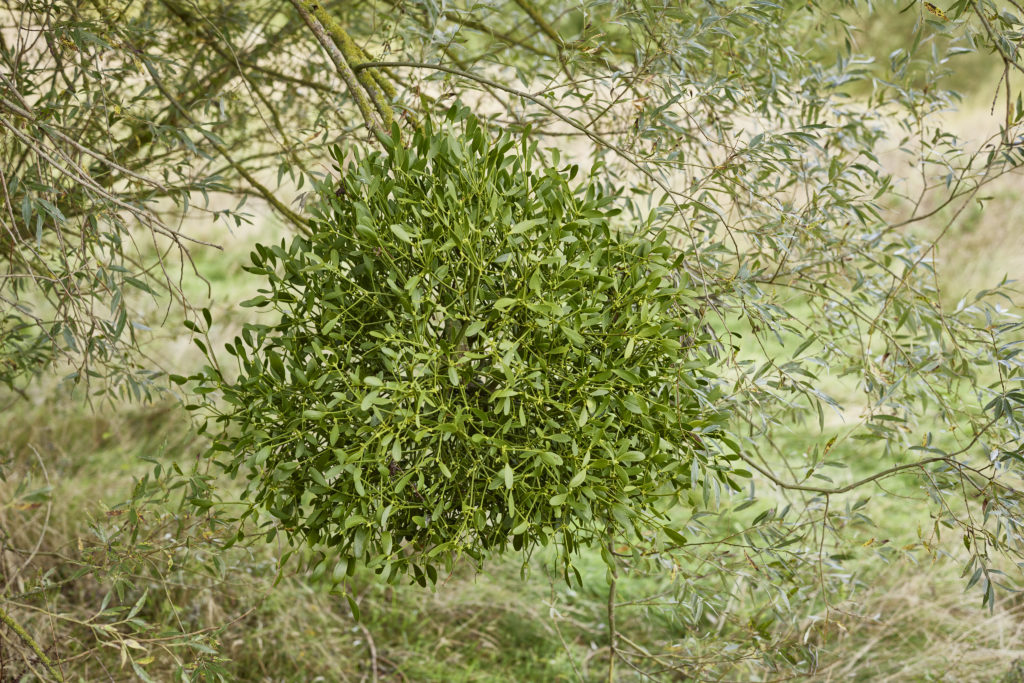

Possibly the single species which has taken up the most DToL time in 2022 is the European mistletoe. Viscum album boasts the largest genome in Britain and Ireland, clocking in at 90 Gbp and 30 times the size of the human genome. Why? Nobody is quite sure, but that’s a question which our reference genome will help answer in future.
Samples from a female mistletoe were collected by the Kew Gardens team in September 2020 in southwest London. The next year, 2021, focused on extracting the plant’s DNA and sequencing its DNA data. This year our bioinformaticians assembled the genome along ten colossal chromosomes. A special mention to Lucia Campos-Dominguez at the University of Edinburgh, who spent the last three months curating the genome – essentially scrolling through, chromosome by chromosome, checking for errors and inversions. Barring some checking of our work and a bit of head-scratching over how to upload this massive genome onto public databases, we can now declare that a reference genome assembly of Britain and Ireland’s biggest genome is now complete!
Read more about the trials and triumphs of assembling the mistletoe genome here.
Protist genomes ready & DNA barcoding successes

Three protist Genome Notes are on the horizon thanks to collaboration between the Earlham Institute, CCAP, the University of Oxford, Marine Biological Association and Sanger Institute. COPO brokered the metadata for the three species, as it has done for all of the DToL samples – now making up a significant proportion of all Earth Biogenome Project standard genomes. A paper on generating annotated genomes from single cells will be published early in 2023.
Earlham’s public engagement initiative, Barcoding the Broads, has trained more than 120 people in DNA barcoding, with four schools in Norfolk and one school in London now conducting independent experiments. Enabling Connections funds supported a number of projects, from a new DNA barcoding hub at the Bayfordbury Field Station in Hertfordshire to work with Kew Gardens and the Norfolk Fungus Study Group. The latter has provided training and resources for community scientists who have identified more than 40 fungi species.
Annotations, data portal features and geocaching

At the start of this year, EMBL-EBI’s DToL team reached the first 100 annotated genomes for new species. This number has steadily increased and these new genome annotations are openly available through the Ensembl DToL page and Ensembl Rapid Release.
The team also made some exciting updates to the DToL Data Portal – an open access platform that pulls together data from across the DToL project making it available all in one place. Users can track the sequencing progress of their species of interest using the Data Portal’s status tracking feature. This feature was updated this year to include detailed status updates to carefully track samples at each step of the process. They also added an interactive sampling map that allows users to identify where species samples have been collected.
On the public engagement side of things, the team designed and implemented a new multilingual activity to engage migrant communities with the science underpinning nature in the local area. This activity – called ‘Into Nature’ – combines geocaching with collectable cards that participants can find and use to learn more about the DToL species.
Coastal collections & marine science explored
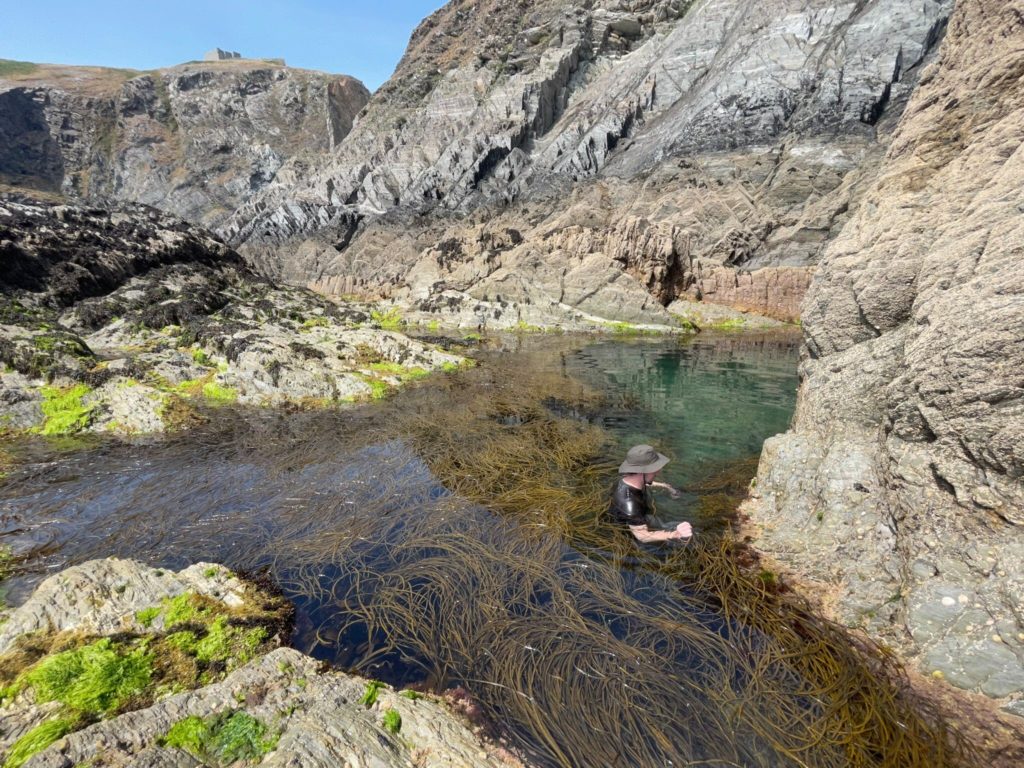

This year the Marine Biological Association collected 1,951 samples representing 274 species, 172 families, and 91 orders. Six species of protists were cultured and successfully harvested at the MBA lab, and 12 species of algae and six species of animals were barcoded in-house. The MBA hosted several visits by experts to collect species, including the Sanger Institute (isopods), Natural History Museum (several groups but with a focus on pericardia), and the University of Bergen (hydroids). Team members received advanced taxonomic training on expert courses for both macroalgae and planktonic species that require a higher level of taxonomic skill for correct identification.
The team attended the Lundy Island Marine Festival where species were sampled and processed in tandem with other DToL parters. The MBA ran a bioblitz which involved both sampling of species from the intertidal zone and educational talks to engage schoolchildren in the project, the wider field of DNA, and the concept of scientific ethics.
Scientific dissemination included an MBA invited keynote presentation given at the Station Biologique de Roscoff’s 150th anniversary, an article in the Marine Biologist international magazine and Biochemist magazine, a Young Marine Biologist careers talk and club event to introduce morphological and molecular identification of species, and genome note publications for marine species that have been sequenced as part of the project. Dissemination to the wider public included a newspaper article in The Times, a Times radio interview, an MBA Twitter takeover week, and seven blogs. Publicly available media made by the MBA team include eleven YouTube videos and a video for the Royal Society Summer Science Exhibition hosted on the DToL web site.
Museum to mountains: epic arthropod collecting
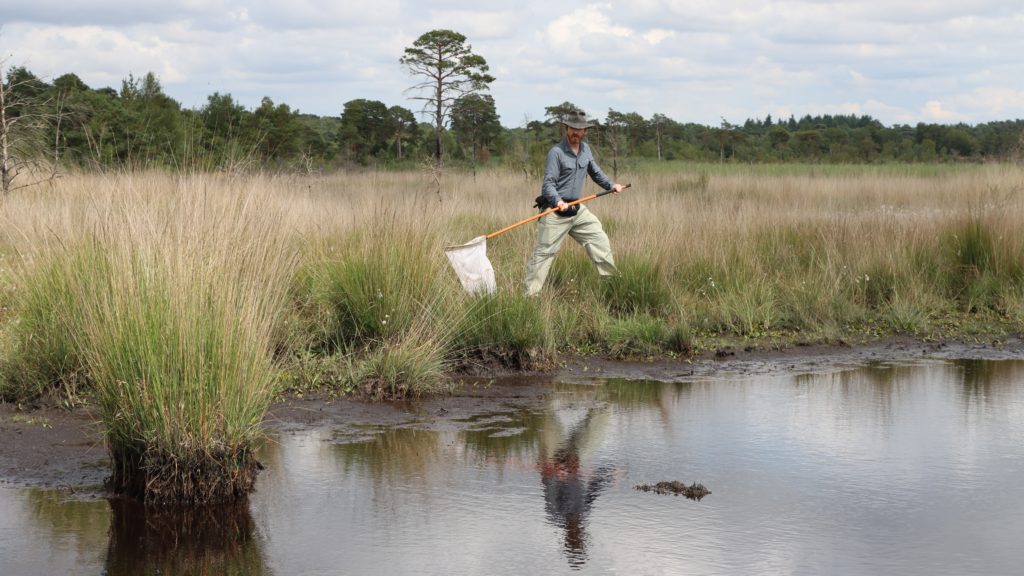

It has been an incredibly successful year of sampling for the Natural History Museum (NHM) DToL team, making up for the 18 “lost COVID months”. Although we had sporadic collecting trips in the first few months of the year, the sampling season truly kicked off in late June (prime ‘beetle season’, as we all know), when the NHM crew teamed up with entomologists from Natural England for a Bioblitz in the Norfolk Broads. Joined by members of the UK Barcode of Life (UKBoL) project, over 550 specimens destined for whole genome or DNA barcoding were collected.
The DToL team then travelled back to Norfolk for the Dipterists’ Forum field meeting in July. There, 561 specimens were identified and frozen, of which 219 were Diptera. A total of 288 species were new to DToL and 119 specimens to UKBoL. Throughout summer the team squeezed in trips to Thursley Common, Knepp Wildwood, Tudeley Woods and Lundy Island.
The sampling season finished off with another summer trip in late August to Beinn Eighe, in the Scottish Highlands. Not only was the NHM team accompanied by members of the National Museums of Scotland, the Highland Biological Recording Group and NatureScot, a Channel 5 film crew were also present, there to film one day of sampling, attempting to capture the frantic behind the scenes of DToL entomological work. Despite the midges and the indecisive weather, there were cracks of sun and team DToL were able to collect 160 species of invertebrates, capping off an incredibly fruitful year of collecting.
A special mention goes to all our external submitters and museum curators who helped with DToL, either by submitting specimens (through individual submissions or through a society), species identification, or through finalising barcode interpretations.
Wytham’s sunny spring & primary school projects
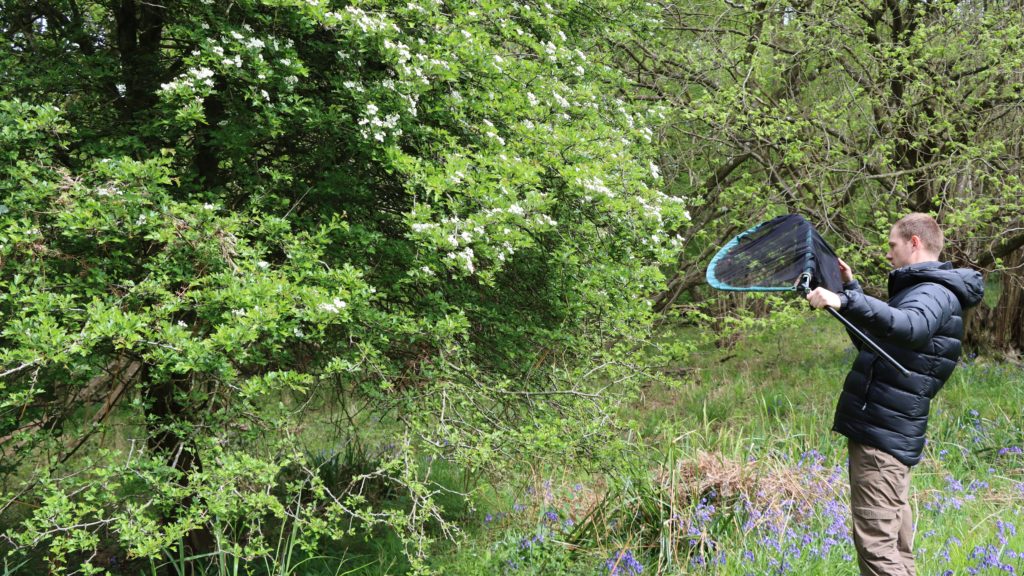

A warm and sunny spring meant that collecting species for genome sequencing at Wytham Woods got off to a rapid start in 2022, although the continued dry weather proved tough for many insects as the hot summer progressed. Activity at the Wytham site included:
- Undergraduate student James McCulloch joining Liam Crowley for collecting and processing specimens during the summer.
- Louis Lofthouse taking on the task of mounting and accessioning voucher specimens.
- Peter Mulhair collecting aquatic taxa and analysing data
- Katie Whale joined us as a Schools Liaison and BugBlitz Coordinator, engaging hundreds of primary school children in the project (more on that here)
This year also saw the establishment of a local volunteer collectors group, involving people with a range of experience from enthusiastic students to national experts.
Among the 750 species collected for genome sequencing in 2022, highlights include:
- the rare Brown Spruce Longhorn Beetle found by a school pupil
- the first Emperor moth from Wytham, a Rugged Oil Beetle and a Sabre Wasp
The team published many Genome Notes, including that of the European badger (Meles meles), and also had two peer-reviewed publications accepted based on analysis of Darwin Tree of Life genome data.
Moving beyond Wytham, we spread our metaphorical net wide to explore satellite sites including a local reed bed site, Withymead, and coordinated with Gloucestershire Wildlife Trust and Royal Entomological Society to collect the rare Large Blue butterfly, a species for which genome data is keenly sought for conservation applications (read more here).
DToL goes to the Royal Society
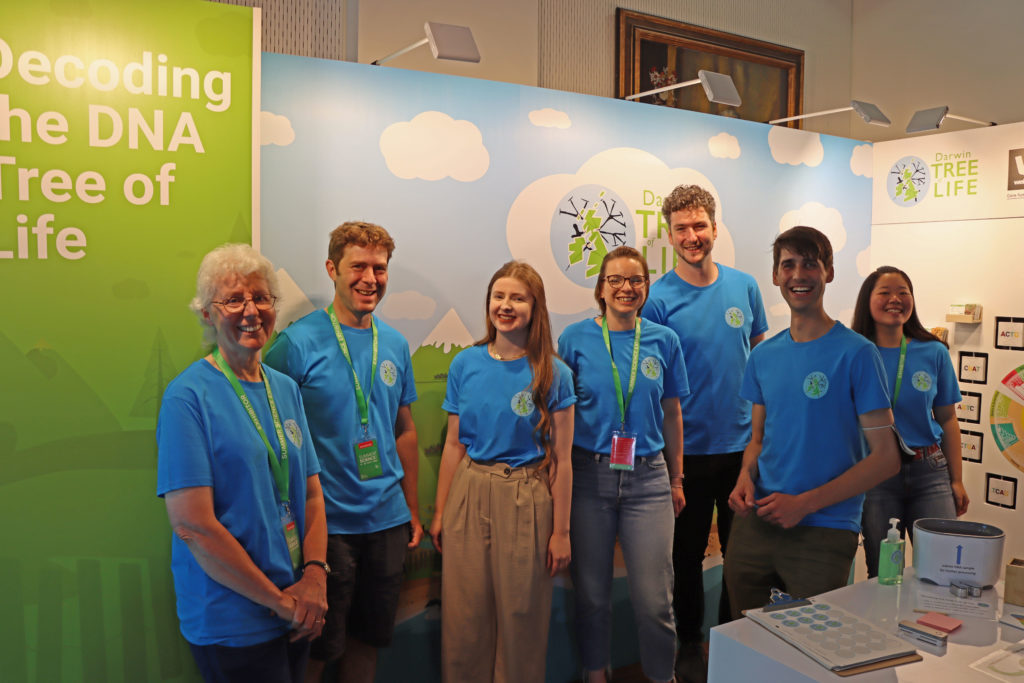

Public engagement with Darwin Tree of Life reached new heights in 2022, with everyone from school children to citizen science groups getting involved, from the Lizard in Cornwall to the Isle of Sky in the Hebrides. Possibly the best part was being invited to exhibit at the Royal Society’s Summer Science Exhibition, a five-day centrepiece of the public science calendar. We think the stats speak for themselves.
- 6,774 visitors, many of whom spoke to the…
- 41 members of the DToL team who travelled to London from partner institutes across the country, and who racked up…
- 360 hours of public engagement between them, and gave out…
- 3,500 fact cards on key species, 2,000 ‘build your own habitat’ postcards, 500 eco-friendly stickers and 800 leaflets, as well as appearing at…
- 3 careers talks with 14-16 years olds from 33 schools.
The verdict? The DToL exhibit was one of the top three most popular on the Royal Society website at the end of the event.
Darwin Tree of Life is a huge collaborative effort spanning the length and breadth of these islands, and only together could we have achieved these fantastic milestones. Here’s to a bright 2023, cracking more genomic challenges, submitting even more species to scientists everywhere, and moving closer to our goal of sequencing everything.